Hi, I need help to check on the code whether is it correct for segmentation as I’m unable to get the bounding boxes and the mask correctly.
import numpy as np
import cv2
from PIL import Image
from hailo_sdk_client import ClientRunner, InferenceContext
import os
# ---------------- CONFIG ----------------
HAR_PATH = "./modelA.har"
IMAGE_PATH = "./test.bmp"
INPUT_SIZE = (416, 416)
NUM_CLASSES = 2
CLASS_NAMES = \["fail", "pass"\]
CONF_THRESH = 0.75
IOU_THRESH = 0.4
MASK_THRESH = 0.5
TOP_K = 1 # only keep top 1 detection
# ---------------- HELPERS ----------------
def sigmoid(x):
return 1 / (1 + np.exp(-x))
def softmax(x, axis=-1):
x = x - np.max(x, axis=axis, keepdims=True)
e = np.exp(x)
return e / (np.sum(e, axis=axis, keepdims=True) + 1e-12)
def compute_iou(box, boxes):
x1 = np.maximum(box\[0\], boxes\[:,0\])
y1 = np.maximum(box\[1\], boxes\[:,1\])
x2 = np.minimum(box\[2\], boxes\[:,2\])
y2 = np.minimum(box\[3\], boxes\[:,3\])
inter = np.maximum(0, x2-x1) \* np.maximum(0, y2-y1)
a1 = (box\[2\]-box\[0\])\*(box\[3\]-box\[1\])
a2 = (boxes\[:,2\]-boxes\[:,0\])\*(boxes\[:,3\]-boxes\[:,1\])
return inter / (a1 + a2 - inter + 1e-12)
def non_max_suppression(boxes, scores, iou_thresh=0.4):
idxs = np.argsort(scores)\[::-1\]
keep=\[\]
while idxs.size>0:
cur = idxs\[0\]
keep.append(cur)
if idxs.size==1:
break
ious = compute_iou(boxes\[cur\], boxes\[idxs\[1:\]\])
idxs = idxs\[1:\]\[ious < iou_thresh\]
return keep
# ---------------- PROCESS ----------------
def preprocess_image(path):
img = Image.open(path).convert("RGB").resize(INPUT_SIZE)
arr = np.array(img).astype(np.float32)/255.0
arr = np.expand_dims(arr, axis=0) # NHWC
return arr, img
def reconstruct_har(outputs, orig_img_size):
"""
Reconstruct boxes and masks from HAR outputs (YOLOX-mask style)
"""
img_w, img_h = orig_img_size
out0, out1, out2 = outputs
# --- proto map ---
proto = np.transpose(out0, (0,3,1,2)) # NHWC -> NCHW
proto_map = proto\[0\] # (mask_dim, H, W)
# --- mask coefficients ---
mask_coeffs = out1\[0,0,:,:\] # (num_anchors, mask_dim)
# --- raw predictions ---
pred_raw = out2\[0,0,:,:\] # (num_anchors, last_dim)
# --- decode cx, cy, w, h ---
cx = sigmoid(pred_raw\[:,0\]) \* INPUT_SIZE\[0\] # already normalized 0-1
cy = sigmoid(pred_raw\[:,1\]) \* INPUT_SIZE\[1\]
# HAR outputs w/h are usually log-space relative to grid cell (YOLOX style)
stride = INPUT_SIZE\[0\] / out0.shape\[1\] # assume square input
w = np.maximum(pred_raw\[:,2\], 1.0) \* stride # ensure minimum size 1
h = np.maximum(pred_raw\[:,3\], 1.0) \* stride
print("cx:", cx.min(), cx.max())
print("cy:", cy.min(), cy.max())
print("w:", w.min(), w.max())
print("h:", h.min(), h.max())
# --- construct boxes ---
x1 = np.clip(cx - w/2, 0, img_w)
y1 = np.clip(cy - h/2, 0, img_h)
x2 = np.clip(cx + w/2, 0, img_w)
y2 = np.clip(cy + h/2, 0, img_h)
boxes = np.stack(\[x1, y1, x2, y2\], axis=1)
# --- class probabilities ---
class_logits = pred_raw\[:, -NUM_CLASSES:\]
probs = softmax(class_logits, axis=1)
class_ids = np.argmax(probs, axis=1)
scores = probs\[np.arange(len(probs)), class_ids\]
# --- filter by confidence ---
keep_mask = scores > CONF_THRESH
boxes = boxes\[keep_mask\]
scores = scores\[keep_mask\]
class_ids = class_ids\[keep_mask\]
mask_coeffs = mask_coeffs\[keep_mask\]
if len(scores) == 0:
print("No detections above threshold.")
return \[\], \[\], \[\], \[\]
# --- NMS ---
keep_idx = non_max_suppression(boxes, scores, IOU_THRESH)
boxes = boxes\[keep_idx\]
scores = scores\[keep_idx\]
class_ids = class_ids\[keep_idx\]
mask_coeffs = mask_coeffs\[keep_idx\]
# --- keep top K detections ---
if len(scores) > TOP_K:
top_idx = np.argsort(scores)\[-TOP_K:\]
boxes = boxes\[top_idx\]
scores = scores\[top_idx\]
class_ids = class_ids\[top_idx\]
mask_coeffs = mask_coeffs\[top_idx\]
# --- reconstruct masks ---
final_masks = \[\]
for coeff in mask_coeffs:
mask = np.tensordot(coeff, proto_map, axes=\[0,0\])
mask = sigmoid(mask)
mask = cv2.resize(mask, (img_w, img_h))
mask = (mask > MASK_THRESH).astype(np.uint8)
mask = cv2.morphologyEx(mask, cv2.MORPH_OPEN, np.ones((3,3), np.uint8))
final_masks.append(mask)
return boxes, scores, class_ids, final_masks
def draw_results(img_pil, boxes, scores, class_ids, masks):
img = np.array(img_pil).copy()
overlay = img.copy()
for i, box in enumerate(boxes.astype(int)):
x1,y1,x2,y2 = box
cls = CLASS_NAMES\[int(class_ids\[i\])\]
score = float(scores\[i\])
color = (0,255,0) if cls=="pass" else (0,0,255)
cv2.rectangle(img, (x1,y1), (x2,y2), color, 2)
cv2.putText(img, f"{cls} {score:.2f}", (x1, max(y1-6,0)), cv2.FONT_HERSHEY_SIMPLEX, 0.5, color, 1)
# overlay mask
if masks:
m = masks\[i\]
colored = np.zeros_like(img)
colored\[:,:,1\] = m\*255 # green mask
alpha = 0.4
overlay = cv2.addWeighted(overlay, 1.0, colored, alpha, 0)
out = cv2.addWeighted(img, 1.0, overlay, 0.4, 0)
out = cv2.cvtColor(out, cv2.COLOR_RGB2BGR)
cv2.imwrite("har_result_wire.png", out)
print("Wrote har_result_wire.png with mask and box.")
# ---------------- MAIN ----------------
def main():
input_data, pil_img = preprocess_image(IMAGE_PATH)
runner = ClientRunner(har=HAR_PATH)
with runner.infer_context(InferenceContext.SDK_NATIVE) as ctx:
outputs = runner.infer(ctx, input_data)
for i, out in enumerate(outputs):
print(f"Output {i}: shape={out.shape}")
boxes, scores, class_ids, masks = reconstruct_har(outputs, pil_img.size\[::-1\])
if len(boxes) > 0:
print("Detections:")
for i in range(len(boxes)):
print(f" - {CLASS_NAMES\[int(class_ids\[i\])\]} {scores\[i\]:.3f} box {boxes\[i\].astype(int).tolist()}")
draw_results(pil_img, boxes, scores, class_ids, masks)
else:
print("No detections after postprocessing.")
if \__name_\_ == "\__main_\_":
main()
I’m not sure is the code incorrect or is the har itself converted from onnx model that affects the segmentation
I convert the onnx model to har using following command
hailo parser onnx modelA.onnx
(I got tried using hailomz but fail to convert it, only this command success convert the model to har)
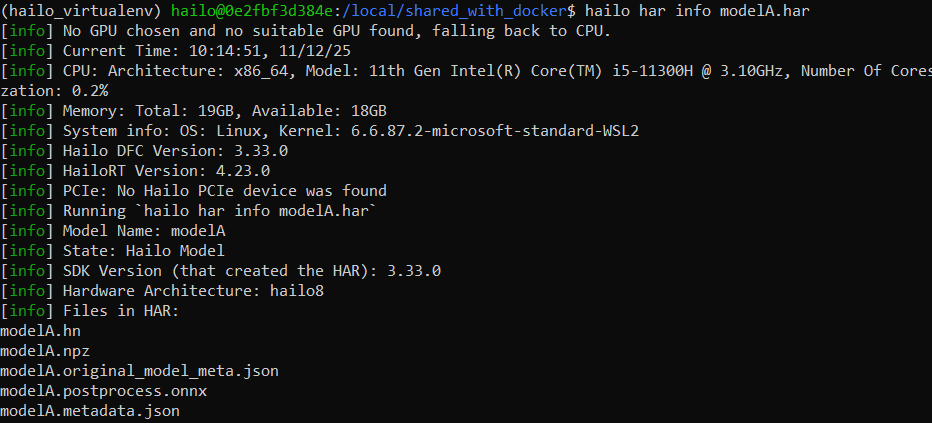
below is about the har info
and i try to do segmentation with the original onnx model with this tutorial and it can segment the images correctly and the following code to test out the segmentation
https://dev.to/andreygermanov/how-to-implement-instance-segmentation-using-yolov8-neural-network-3if9#join_masks
import onnxruntime as ort
import numpy as np
from PIL import Image
import cv2
import matplotlib.pyplot as plt
yolo_classes = \["fail", "pass"\]
# --- Helper functions ---
def sigmoid(z):
return 1 / (1 + np.exp(-z))
def get_mask(row, box, img_width, img_height):
mask = row.reshape(104,104)
mask = sigmoid(mask)
mask = (mask > 0.5).astype('uint8')\*255
x1, y1, x2, y2 = box
mask_x1 = round(x1 / img_width \* 104)
mask_y1 = round(y1 / img_height \* 104)
mask_x2 = round(x2 / img_width \* 104)
mask_y2 = round(y2 / img_height \* 104)
mask = mask\[mask_y1:mask_y2, mask_x1:mask_x2\]
img_mask = Image.fromarray(mask, "L")
img_mask = img_mask.resize((round(x2-x1), round(y2-y1)))
return np.array(img_mask)
def get_polygons(mask):
contours, \_ = cv2.findContours(mask, cv2.RETR_EXTERNAL, cv2.CHAIN_APPROX_SIMPLE)
polygons = \[\]
for contour in contours:
contour = contour.reshape(-1,2)
polygons.append(contour)
return polygons
def intersection(box1, box2):
x1 = max(box1\[0\], box2\[0\])
y1 = max(box1\[1\], box2\[1\])
x2 = min(box1\[2\], box2\[2\])
y2 = min(box1\[3\], box2\[3\])
return max(0, x2-x1) \* max(0, y2-y1)
def union(box1, box2):
area1 = (box1\[2\]-box1\[0\])\*(box1\[3\]-box1\[1\])
area2 = (box2\[2\]-box2\[0\])\*(box2\[3\]-box2\[1\])
return area1 + area2 - intersection(box1, box2)
def iou(box1, box2):
return intersection(box1, box2) / union(box1, box2)
# --- Run model ---
def run_model(input):
model = ort.InferenceSession("modelA.onnx")
outputs = model.run(None, {"images": input})
return outputs
# --- Process outputs ---
def process_output(outputs, img_width, img_height):
output0 = outputs\[0\]\[0\].transpose()
output1 = outputs\[1\]\[0\].astype("float")
boxes = output0\[:, 0:6\]
masks = output0\[:, 6:\]
output1 = output1.reshape(32, 104\*104)
masks = masks @ output1
boxes = np.hstack((boxes, masks))
objects = \[\]
for row in boxes:
prob = row\[4:6\].max()
if prob < 0.5:
continue
class_id = row\[4:6\].argmax()
label = yolo_classes\[class_id\]
xc, yc, w, h = row\[:4\]
x1 = (xc - w/2)/416 \* img_width
y1 = (yc - h/2)/416 \* img_height
x2 = (xc + w/2)/416 \* img_width
y2 = (yc + h/2)/416 \* img_height
mask = get_mask(row\[6:\], (x1, y1, x2, y2), img_width, img_height)
polygons = get_polygons(mask)
objects.append(\[x1, y1, x2, y2, label, prob, mask, polygons\])
# Non-max suppression
objects.sort(key=lambda x: x\[5\], reverse=True)
result = \[\]
while objects:
result.append(objects\[0\])
objects = \[obj for obj in objects\[1:\] if iou(obj, result\[-1\]) < 0.5\]
return result
# --- Load image ---
img = Image.open("test.bmp").convert("RGB")
img_width, img_height = img.size
img_resized = img.resize((416,416))
input_array = np.array(img_resized).transpose(2,0,1).reshape(1,3,416,416).astype('float32')/255.0
# --- Run model & process output ---
outputs = run_model(input_array)
results = process_output(outputs, img_width, img_height)
for i, out in enumerate(outputs):
print(f"ONNX output{i} shape={out.shape}, min={out.min()}, max={out.max()}, mean={out.mean():.4f}")
# Save for comparison
import os
os.makedirs("onnx_diagnostics", exist_ok=True)
for i, out in enumerate(outputs):
np.save(f"onnx_diagnostics/output{i}.npy", out)
# --- Draw results ---
img_np = np.array(img)
for x1, y1, x2, y2, label, prob, mask, polygons in results:
\# overlay mask
for poly in polygons:
cv2.drawContours(img_np\[int(y1):int(y2), int(x1):int(x2)\], \[np.array(poly)\], -1, (0,255,0), -1)
\# draw bounding box
cv2.rectangle(img_np, (int(x1), int(y1)), (int(x2), int(y2)), (255,0,0), 2)
\# label
cv2.putText(img_np, label, (int(x1), int(y1)-5), cv2.FONT_HERSHEY_SIMPLEX, 0.7, (255,255,255), 2)
# --- Show image ---
plt.figure(figsize=(10,10))
plt.imshow(img_np)
plt.axis('off')
plt.show()
cv2.imshow("Segmentation Result", img_np)
cv2.waitKey(0)
cv2.destroyAllWindows()
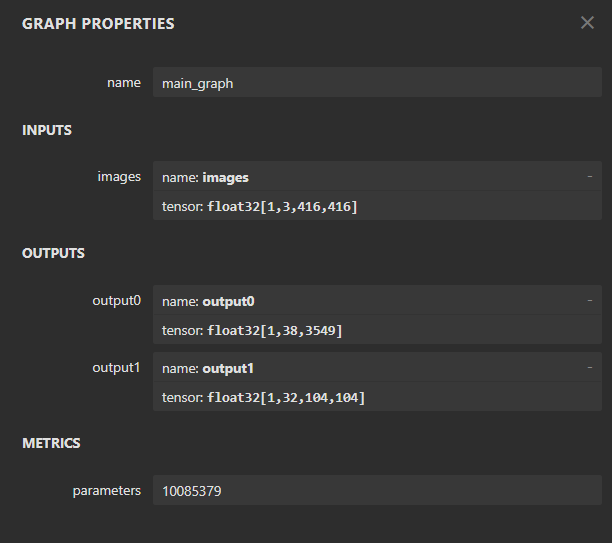
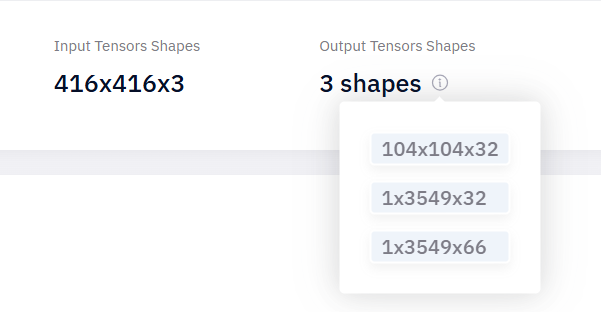
also have question is it normal that it will change the output shape when it convert from onnx to har
the original onnx model has this shape
after convert it to har, it becomes 3 output shape
is it because the output shape has changed that cause the segmentation failed
(something like it affect the boxes channel? (cx, cy, w, h))
please help me i dont now what i do, much appreciate ![]()