Hi @nina-vilela ,
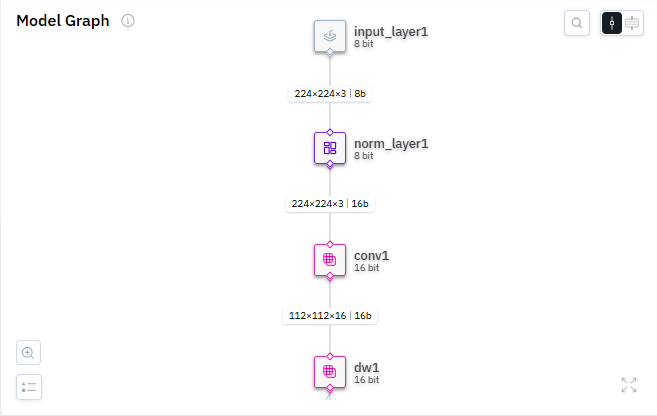
Thank you for the reply. To answer your first question, none of these layers are output layers or input ones for that matter. The nearest layer to the input layer is conv1 and its third. Check this screenshot below:
Yes, I can share my evaluation code for the compiled model. It is a little huge, but most of it is to get the true labels from the csv file and validate against the predicted ones. While creating bindings for each image, I included metadata that has the filename, true classes for the respective attributes of the image. So that in my inference callback, I am able to validate my model prediction to the true labels and calculate overall and joint accuracy.
To make a quick note, Testing this inference pipeline with a few of my test images and also the AlexNet model, which compiled successfully, It works perfectly fine in its validation process.
#!/usr/bin/env python3
“”"
Refined Hailo-8 validation and inference pipeline
Multihead output
GTSRB head
TypeSignal head
CSV labels expected columns
name of file
gtrsb
gtrsb_name
type_signal
type_signal_name
Notes
The script uses gtrsb_name and type_signal_name from the CSV for readable labels
Accuracy is computed for each head and also joint accuracy
Optionally prints a predicted vs actual report for images in a separate folder
“”"
from future import annotations
import argparse
import logging
import os
import threading
import time
from concurrent.futures import ThreadPoolExecutor, Future
from dataclasses import dataclass
from typing import Dict, List, Optional, Tuple
import cv2
import numpy as np
import pandas as pd
from HailoInferClass import HailoInfer
@dataclass
class InferenceConfig:
hef: str
images_dir: str
csv_labels: Optional[str] = None
output_dir: Optional[str] = None
batch_size: int = 24
num_workers: int = 4
num_images: Optional[int] = None
log_level: str = "INFO"
verbose: bool = False
throughput_mode: str = "end_to_end"
device_id: int = 0
gtsrb_head_key_hint: str = "gtsrb"
type_head_key_hint: str = "type"
report_dir: Optional[str] = None
report_samples: int = 10
def preprocess_image_uint8(image_path: str, input_shape: Tuple[int, int, int]) → np.ndarray:
img = cv2.imread(image_path)
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
img = cv2.resize(img, (input_shape[1], input_shape[0]))
return img.astype(np.uint8)
def _safe_int(x) → Optional[int]:
if pd.isna(x):
return None
try:
return int(x)
except Exception:
return None
def load_labels_from_csv_multihead(
csv_path: str,
) → Tuple[Dict[str, Dict[str, Optional[int]]], Dict[int, str], Dict[int, str]]:
df = pd.read_csv(csv_path)
required = {"name of file", "gtrsb", "gtrsb_name", "type_signal", "type_signal_name"}
missing = required.difference(set(df.columns))
if missing:
raise ValueError(f"CSV missing columns: {sorted(list(missing))}")
labels_map: Dict[str, Dict[str, Optional[int]]] = {}
gtsrb_names: Dict[int, str] = {}
type_names: Dict[int, str] = {}
for _, row in df.iterrows():
fname = str(row["name of file"])
g_id = _safe_int(row["gtrsb"])
t_id = _safe_int(row["type_signal"])
labels_map[fname] = {"gtsrb": g_id, "type": t_id}
if g_id is not None and g_id not in gtsrb_names:
g_name_val = row["gtrsb_name"]
gtsrb_names[g_id] = str(g_name_val) if not pd.isna(g_name_val) else f"Class {g_id}"
if t_id is not None and t_id not in type_names:
t_name_val = row["type_signal_name"]
type_names[t_id] = str(t_name_val) if not pd.isna(t_name_val) else f"Class {t_id}"
return labels_map, gtsrb_names, type_names
class HailoApp:
def __init__(self, config: InferenceConfig) -> None:
self.config = config
self.model: Optional[HailoInfer] = None
self.input_shape: Optional[Tuple[int, int, int]] = None
if not os.path.isdir(self.config.images_dir):
raise FileNotFoundError(f"Images directory not found: {self.config.images_dir}")
all_files = sorted(os.listdir(self.config.images_dir))
self.image_names = [f for f in all_files if f.lower().endswith((".jpg", ".jpeg", ".png"))]
self.image_files = [os.path.join(self.config.images_dir, f) for f in self.image_names]
self.true_labels_map: Dict[str, Dict[str, Optional[int]]] = {}
self.gtsrb_names: Dict[int, str] = {}
self.type_names: Dict[int, str] = {}
if self.config.csv_labels:
if os.path.exists(self.config.csv_labels):
self.true_labels_map, self.gtsrb_names, self.type_names = load_labels_from_csv_multihead(
self.config.csv_labels
)
else:
logging.warning(
"CSV labels path provided but not found: %s. Continuing without CSV labels.",
self.config.csv_labels,
)
self.config.csv_labels = None
self.filenames_for_hailo: Dict[int, List] = {}
for i, image_name in enumerate(self.image_names):
targets = self.true_labels_map.get(image_name, {})
self.filenames_for_hailo[i] = [
image_name,
{"gtsrb": targets.get("gtsrb"), "type": targets.get("type")},
]
self._lock = threading.Lock()
self.correct_gtsrb = 0
self.correct_type = 0
self.correct_joint = 0
self.total_samples = 0
self.batch_acc_gtsrb: List[float] = []
self.batch_acc_type: List[float] = []
self.batch_acc_joint: List[float] = []
self.output_names: List[str] = []
self.report_records: List[Dict[str, object]] = []
self.report_name_set: Optional[set] = None
if self.config.report_dir:
self.report_name_set = self._collect_report_names(self.config.report_dir)
def _collect_report_names(self, report_dir: str) -> set:
if not os.path.isdir(report_dir):
raise FileNotFoundError(f"Report directory not found: {report_dir}")
files = sorted(os.listdir(report_dir))
names = [f for f in files if f.lower().endswith((".jpg", ".jpeg", ".png"))]
return set(names[: max(0, int(self.config.report_samples))])
def load_model(self) -> None:
self.model = HailoInfer(
hef_path=self.config.hef,
batch_size=self.config.batch_size,
filenames=self.filenames_for_hailo,
input_type="UINT8",
output_type="UINT8",
)
self.input_shape = self.model.get_input_shape()
self.output_names = [
"alexnet_third_version/conv8",#globalavgpool_mobinet_patched/fc19" gtrsb alexnet_g86_t98_conv_instead_of_linear_dynamic_batch/conv8 globalavgpool_mobinet_patched_dynamic_axes/fc19
"alexnet_third_version/conv9",#globalavgpool_mobinet_patched/fc20" type alexnet_g86_t98_conv_instead_of_linear_dynamic_batch/conv9 globalavgpool_mobinet_patched_dynamic_axes/fc20
]
logging.info("Model input shape: %s", str(self.input_shape))
def close(self) -> None:
if self.model:
try:
self.model.close()
except Exception as e:
logging.warning("Exception while closing model: %s", e)
finally:
logging.info("Hailo device released.")
def _image_batches(self) -> List[List[str]]:
if not self.image_files:
return []
num_to_run = self.config.num_images if self.config.num_images is not None else len(self.image_files)
num_to_run = min(num_to_run, len(self.image_files))
files = self.image_files[:num_to_run]
batches: List[List[str]] = []
for i in range(0, len(files), self.config.batch_size):
batches.append(files[i : i + self.config.batch_size])
return batches
def prepare_batch(self, batch_files: List[str]) -> List[np.ndarray]:
assert self.input_shape is not None
frames: List[np.ndarray] = []
for p in batch_files:
try:
frames.append(preprocess_image_uint8(p, self.input_shape))
except Exception as e:
logging.warning("Failed to preprocess %s: %s", p, e)
return frames
def _extract_targets_from_meta(self, meta: dict) -> Tuple[Optional[int], Optional[int], Optional[str]]:
idx = meta.get("index")
if idx is None:
return None, None, None
try:
idx_int = int(idx)
except Exception:
return None, None, None
entry = self.filenames_for_hailo.get(idx_int)
if not entry:
return None, None, None
filename = entry[0]
targets = entry[1] if len(entry) > 1 and isinstance(entry[1], dict) else {}
true_gtsrb = targets.get("gtsrb")
true_type = targets.get("type")
return true_gtsrb, true_type, filename
def _softmax_probs(self, logits_1d: np.ndarray) -> np.ndarray:
x = logits_1d.astype(np.float32)
x = x - np.max(x)
exp_scores = np.exp(x)
return exp_scores / np.sum(exp_scores)
def _head_from_named_buffers(self, named: Dict[str, np.ndarray]) -> Tuple[np.ndarray, np.ndarray]:
g_key = self.output_names[0] #"globalavgpool_mobinet_patched/fc19" #"alexnet_g86_t98_conv_instead_of_linear_dynamic_batch/conv8"
t_key = self.output_names[1] #"globalavgpool_mobinet_patched/fc20" #"alexnet_g86_t98_conv_instead_of_linear_dynamic_batch/conv9"
if g_key not in named or t_key not in named:
raise RuntimeError(f"Missing expected output names. Got keys: {sorted(list(named.keys()))}")
gtsrb_logits = np.asarray(named[g_key]).reshape(-1)
type_logits = np.asarray(named[t_key]).reshape(-1)
return gtsrb_logits, type_logits
def _maybe_store_report_row(
self,
filename: Optional[str],
true_gtsrb_id: Optional[int],
true_type_id: Optional[int],
pred_gtsrb_id: int,
pred_type_id: int,
) -> None:
if not self.report_name_set:
return
if not filename:
return
if filename not in self.report_name_set:
return
if len(self.report_records) >= int(self.config.report_samples):
return
pred_gtsrb_name = self.gtsrb_names.get(pred_gtsrb_id, f"Class {pred_gtsrb_id}")
pred_type_name = self.type_names.get(pred_type_id, f"Class {pred_type_id}")
true_gtsrb_name = self.gtsrb_names.get(true_gtsrb_id, f"Class {true_gtsrb_id}") if true_gtsrb_id is not None else "None"
true_type_name = self.type_names.get(true_type_id, f"Class {true_type_id}") if true_type_id is not None else "None"
row = {
"file": filename,
"true_gtsrb_id": true_gtsrb_id,
"true_gtsrb_name": true_gtsrb_name,
"pred_gtsrb_id": pred_gtsrb_id,
"pred_gtsrb_name": pred_gtsrb_name,
"gtsrb_correct": (true_gtsrb_id is not None and pred_gtsrb_id == int(true_gtsrb_id)),
"true_type_id": true_type_id,
"true_type_name": true_type_name,
"pred_type_id": pred_type_id,
"pred_type_name": pred_type_name,
"type_correct": (true_type_id is not None and pred_type_id == int(true_type_id)),
}
row["joint_correct"] = bool(row["gtsrb_correct"] and row["type_correct"])
self.report_records.append(row)
def inference_callback(self, infer_results=None, bindings_list=None, validate: bool = True, **kwargs) -> None:
if bindings_list is None:
logging.warning("No bindings_list received in callback.")
return
batch_true_gtsrb: List[int] = []
batch_true_type: List[int] = []
batch_pred_gtsrb: List[int] = []
batch_pred_type: List[int] = []
for info in bindings_list:
binding = info.get("binding")
meta = info.get("metadata", {})
try:
named_outputs = {name: binding.output(name).get_buffer() for name in self.output_names}
except Exception as e:
logging.warning("Failed to get output buffer: %s", e)
continue
try:
gtsrb_logits, type_logits = self._head_from_named_buffers(named_outputs)
except Exception as e:
logging.warning("Failed to parse multihead outputs: %s", e)
continue
# No softmax for UINT8 outputs
# Argmax of logits equals argmax of softmax logits
pred_gtsrb_id = int(np.argmax(gtsrb_logits))
pred_type_id = int(np.argmax(type_logits))
if self.total_samples == 0:
logging.info("gtsrb out shape %s dtype %s", gtsrb_logits.shape, gtsrb_logits.dtype)
logging.info("type out shape %s dtype %s", type_logits.shape, type_logits.dtype)
logging.info("gtsrb first10 %s", gtsrb_logits[:10])
logging.info("type first10 %s", type_logits[:10])
true_gtsrb_id, true_type_id, filename = self._extract_targets_from_meta(meta)
self._maybe_store_report_row(filename, true_gtsrb_id, true_type_id, pred_gtsrb_id, pred_type_id)
if validate and true_gtsrb_id is not None and true_type_id is not None:
batch_true_gtsrb.append(int(true_gtsrb_id))
batch_true_type.append(int(true_type_id))
batch_pred_gtsrb.append(int(pred_gtsrb_id))
batch_pred_type.append(int(pred_type_id))
if validate and batch_true_gtsrb:
tg = np.array(batch_true_gtsrb, dtype=np.int32)
tt = np.array(batch_true_type, dtype=np.int32)
pg = np.array(batch_pred_gtsrb, dtype=np.int32)
pt = np.array(batch_pred_type, dtype=np.int32)
acc_g = float(np.mean(pg == tg))
acc_t = float(np.mean(pt == tt))
acc_joint = float(np.mean((pg == tg) & (pt == tt)))
with self._lock:
self.correct_gtsrb += int(np.sum(pg == tg))
self.correct_type += int(np.sum(pt == tt))
self.correct_joint += int(np.sum((pg == tg) & (pt == tt)))
self.total_samples += int(len(tg))
self.batch_acc_gtsrb.append(acc_g)
self.batch_acc_type.append(acc_t)
self.batch_acc_joint.append(acc_joint)
logging.info(
"Batch processed: n=%d acc_gtsrb=%.2f%% acc_type=%.2f%% acc_joint=%.2f%%",
len(tg),
acc_g * 100.0,
acc_t * 100.0,
acc_joint * 100.0,
)
def run(self) -> None:
if not self.image_files:
logging.error("No images found in: %s", self.config.images_dir)
return
if self.model is None or self.input_shape is None:
self.load_model()
batches = self._image_batches()
if not batches:
logging.error("No batches created from images.")
return
if self.config.output_dir:
os.makedirs(self.config.output_dir, exist_ok=True)
if self.config.throughput_mode == "end_to_end":
self._end_to_end_start = time.perf_counter()
with ThreadPoolExecutor(max_workers=max(1, self.config.num_workers)) as executor:
preprocess_futures: List[Tuple[int, Future]] = []
next_batch_idx = 0
first_run_async_called = False
if next_batch_idx < len(batches):
fut = executor.submit(self.prepare_batch, batches[next_batch_idx])
preprocess_futures.append((next_batch_idx, fut))
next_batch_idx += 1
self._host_start = time.perf_counter()
while preprocess_futures:
idx, future = preprocess_futures.pop(0)
frames = future.result()
if frames:
if not first_run_async_called:
self._device_start = time.perf_counter()
first_run_async_called = True
assert self.model is not None
try:
self.model.run_async(frames, self.inference_callback)
except Exception as e:
logging.error("Error when calling model.run_async: %s", e)
if next_batch_idx < len(batches):
fut = executor.submit(self.prepare_batch, batches[next_batch_idx])
preprocess_futures.append((next_batch_idx, fut))
next_batch_idx += 1
self._host_end = time.perf_counter()
if self.model and getattr(self.model, "last_infer_job", None):
last_job = self.model.last_infer_job
try:
if hasattr(last_job, "wait"):
last_job.wait(1000)
else:
time.sleep(0.1)
except Exception as e:
logging.warning("Exception while waiting for last_infer_job: %s", e)
self.close()
self._report_metrics()
self._print_report_rows()
def _print_report_rows(self) -> None:
if not self.report_records:
return
print("")
print("Predicted vs Actual sample report")
print(f"Report rows: {len(self.report_records)}")
for r in self.report_records:
print("")
print(f"File: {r['file']}")
print(f"GTSRB true: {r['true_gtsrb_name']} pred: {r['pred_gtsrb_name']} correct: {r['gtsrb_correct']}")
print(f"Type true: {r['true_type_name']} pred: {r['pred_type_name']} correct: {r['type_correct']}")
print(f"Joint correct: {r['joint_correct']}")
def _report_metrics(self) -> None:
print(f"Total labeled samples processed: {self.total_samples}")
if self.total_samples > 0:
acc_g = (self.correct_gtsrb / self.total_samples) * 100.0
acc_t = (self.correct_type / self.total_samples) * 100.0
acc_j = (self.correct_joint / self.total_samples) * 100.0
print(f"GTSRB accuracy: {acc_g:.2f} %")
print(f"TypeSignal accuracy: {acc_t:.2f} %")
print(f"Joint accuracy: {acc_j:.2f} %")
else:
print("No labeled samples were processed for accuracy.")
if self.batch_acc_gtsrb:
print(f"Avg batch GTSRB accuracy: {float(np.mean(self.batch_acc_gtsrb)) * 100.0:.2f} %")
if self.batch_acc_type:
print(f"Avg batch TypeSignal accuracy: {float(np.mean(self.batch_acc_type)) * 100.0:.2f} %")
if self.batch_acc_joint:
print(f"Avg batch Joint accuracy: {float(np.mean(self.batch_acc_joint)) * 100.0:.2f} %")
def parse_args() → InferenceConfig:
p = argparse.ArgumentParser(description="Refined Hailo-8 inference pipeline, multihead")
p.add_argument("--hef", required=True, help="Path to compiled Hailo HEF model.")
p.add_argument("--images-dir", required=True, help="Directory with input images.")
p.add_argument("--csv-labels", required=False, default=None, help="CSV with labels and names.")
p.add_argument("--output-dir", required=False, default=None, help="Optional output directory.")
p.add_argument("--batch-size", required=False, type=int, default=24)
p.add_argument("--num-workers", required=False, type=int, default=4)
p.add_argument("--num-images", required=False, type=int, default=None)
p.add_argument("--log-level", required=False, default="INFO")
p.add_argument("--verbose", action="store_true")
p.add_argument("--throughput-mode", required=False, default="end_to_end", choices=["device", "host", "end_to_end"])
p.add_argument("--gtsrb-head-key-hint", required=False, default="gtsrb")
p.add_argument("--type-head-key-hint", required=False, default="type")
p.add_argument("--report-dir", required=False, default=None, help="Folder with images to print predicted vs actual for")
p.add_argument("--report-samples", required=False, type=int, default=10, help="How many report rows to print")
args = p.parse_args()
return InferenceConfig(
hef=args.hef,
images_dir=args.images_dir,
csv_labels=args.csv_labels,
output_dir=args.output_dir,
batch_size=args.batch_size,
num_workers=args.num_workers,
num_images=args.num_images,
log_level=args.log_level,
verbose=args.verbose,
throughput_mode=args.throughput_mode,
gtsrb_head_key_hint=args.gtsrb_head_key_hint,
type_head_key_hint=args.type_head_key_hint,
report_dir=args.report_dir,
report_samples=args.report_samples,
)
def main() → None:
config = parse_args()
numeric_level = getattr(logging, config.log_level.upper(), logging.INFO)
logging.basicConfig(level=numeric_level, format="%(asctime)s [%(levelname)s] %(message)s")
if not os.path.exists(config.hef):
logging.error("HEF model not found: %s", config.hef)
return
try:
app = HailoApp(config)
except Exception as e:
logging.error("Failed to initialize HailoApp: %s", e)
return
try:
app.run()
finally:
app.close()
if name == “main”:
main()
The HailoInferClass.py that acts like the API to communicate with Hailo:
from typing import Tuple, Dict, List
from typing import Callable, Optional
from functools import partial
import numpy as np
import time
from hailo_platform import (HEF, VDevice,FormatType, HailoSchedulingAlgorithm)
from hailo_platform.pyhailort.pyhailort import FormatOrder
class HailoInfer:
def __init__(
self, hef_path: str, batch_size: int = 1,
input_type: Optional[str] = None,
output_type: Optional[str] = None,
filenames:Optional[Dict] = None,
priority: Optional[int] = 0) -> None:
"""
Initialize the HailoAsyncInference class to perform asynchronous inference using a Hailo HEF model.
Args:
hef_path (str): Path to the HEF model file.
batch_size (optional[int]): Number of inputs processed per inference. Defaults to 1.
input_type (Optional[str], optional): Input data type format. Common values: 'UINT8', 'UINT16', 'FLOAT32'.
output_type (Optional[str], optional): Output data type format. Common values: 'UINT8', 'UINT16', 'FLOAT32'.
priority (optional[int]): Scheduler priority value for the model within the shared VDevice context. Defaults to 0.
"""
self.filenames = filenames
params = VDevice.create_params()
# Set the scheduling algorithm to round-robin to activate the scheduler
params.scheduling_algorithm = HailoSchedulingAlgorithm.ROUND_ROBIN
params.group_id = "SHARED"
vDevice = VDevice(params)
self.target = vDevice
self.hef = HEF(hef_path)
ins = self.hef.get_input_vstream_infos()
print("INPUT VSTREAM INFO")
for v in ins:
print("name:", v.name)
print("shape:", v.shape)
print("format type:", v.format.type)
print("format order:", v.format.order)
self.infer_model = self.target.create_infer_model(hef_path)
self.infer_model.set_batch_size(batch_size)
self._set_input_type(input_type)
self._set_output_type(output_type)
self.config_ctx = self.infer_model.configure()
self.configured_model = self.config_ctx.__enter__()
self.configured_model.set_scheduler_priority(priority)
self.last_infer_job = None
self.count_index = 0
def _set_input_type(self, input_type: Optional[str] = None) -> None:
"""
Set the input type for the HEF model. If the model has multiple inputs,
it will set the same type of all of them.
Args:
input_type (Optional[str]): Format type of the input stream.
"""
if input_type is not None:
self.infer_model.input().set_format_type(getattr(FormatType, input_type))
def _set_output_type(self, output_type: Optional[str] = None) -> None:
"""
Set the output type for each model output.
Args:
output_type (Optional[str]): Desired output data type. Common values:
'UINT8', 'UINT16', 'FLOAT32'.
"""
self.nms_postprocess_enabled = False
# If the model uses HAILO_NMS_WITH_BYTE_MASK format (e.g.,instance segmentation),
if self.infer_model.outputs[0].format.order == FormatOrder.HAILO_NMS_WITH_BYTE_MASK:
# Use UINT8 and skip setting output formats
self.nms_postprocess_enabled = True
self.output_type = self._output_data_type2dict("UINT8")
return
# Otherwise, set the format type based on the provided output_type argument
self.output_type = self._output_data_type2dict(output_type)
# Apply format to each output layer
for name, dtype in self.output_type.items():
self.infer_model.output(name).set_format_type(getattr(FormatType, dtype))
def get_vstream_info(self) -> Tuple[list, list]:
"""
Get information about input and output stream layers.
Returns:
Tuple[list, list]: List of input stream layer information, List of
output stream layer information.
"""
return (
self.hef.get_input_vstream_infos(),
self.hef.get_output_vstream_infos()
)
def get_hef(self) -> HEF:
"""
Get a HEF instance
Returns:
HEF: A HEF (Hailo Executable File) containing the model.
"""
return self.hef
def get_input_shape(self) -> Tuple[int, ...]:
"""
Get the shape of the model's input layer.
Returns:
Tuple[int, ...]: Shape of the model's input layer.
"""
return self.hef.get_input_vstream_infos()[0].shape # Assumes one input
def run_async(self, input_batch: List[np.ndarray], inference_callback_fn) -> object:
"""
Run an asynchronous inference job on a batch of preprocessed inputs.
This method reuses a preconfigured model (no reconfiguration overhead),
prepares input/output bindings, launches async inference, and returns
the job handle so that the caller can wait on it if needed.
Args:
input_batch (List[np.ndarray]): A batch of preprocessed model inputs.
inference_callback_fn (Callable): Function to be invoked when inference is complete.
It receives `bindings_list` and additional context.
Returns:
None
"""
start = time.time()
bindings_list = self.create_bindings(input_batch,self.configured_model)
print(f'We are at index:{self.count_index}')
self.configured_model.wait_for_async_ready(timeout_ms=10000)
hailo_bindings = [b['binding'] for b in bindings_list]
# Launch async inference and attach the result handler
self.last_infer_job = self.configured_model.run_async(
hailo_bindings,
partial(inference_callback_fn, bindings_list=bindings_list)
)
print(f"[INFO] Submitted async job handle: {self.last_infer_job}")
if self.last_infer_job is not None:
print("[DEBUG] Waiting for first async batch to complete...")
self.last_infer_job.wait(1000)
print("[INFO] Async batch finished. Device ready for next set.")
def create_bindings(self,input_batch,configured_model):
bindings_list = []
for i,frame in enumerate(input_batch):
output_buffers = {
name:np.empty(
self.infer_model.output(name).shape,
dtype=(getattr(np,self.output_type[name].lower()))
)
for name in self.output_type
}
binding = configured_model.create_bindings(output_buffers=output_buffers)
binding.input().set_buffer(np.array(frame))
current_index = self.count_index
meta = {
"index":current_index,
"filename":self.filenames[current_index][0] if self.filenames else None,
"true class":self.filenames[current_index][1] if self.filenames else None,
"timestamp":time.time()
}
bindings_list.append({'binding':binding,'metadata':meta})
self.count_index+=1
return bindings_list
def is_nms_postprocess_enabled(self) -> bool:
"""
Returns True if the HEF model includes an NMS postprocess node.
"""
return self.nms_postprocess_enabled
def _output_data_type2dict(self, data_type: Optional[str]) -> Dict[str, str]:
"""
Generate a dictionary mapping each output layer name to its corresponding
data type. If no data type is provided, use the type defined in the HEF.
Args:
data_type (Optional[str]): The desired data type for all output layers.
Valid values: 'float32', 'uint8', 'uint16'.
If None, uses types from the HEF metadata.
Returns:
Dict[str, str]: A dictionary mapping output layer names to data types.
"""
valid_types = {"float32", "uint8", "uint16"}
data_type_dict = {}
for output_info in self.hef.get_output_vstream_infos():
name = output_info.name
if data_type is None:
# Extract type from HEF metadata
hef_type = str(output_info.format.type).split(".")[-1]
data_type_dict[name] = hef_type
else:
if data_type.lower() not in valid_types:
raise ValueError(f"Invalid data_type: {data_type}. Must be one of {valid_types}")
data_type_dict[name] = data_type
return data_type_dict
def close(self):
# Wait for the final job to complete before exiting
if self.last_infer_job is not None:
self.last_infer_job.wait(10000)
if self.config_ctx:
self.config_ctx.__exit__(None, None, None)